CAPRI Assessment results
Round 61
Round 61 features four targets, each a different antibody binding to the same antigen. Full assessment details can be found in this file.
Top performers of this Round:
| Predictor | Top-1 | Top-5 | Top-10 |
|---|---|---|---|
| CLUSPRO | 2/1***/1** | 2/1***/1** | 2/1***/1** |
| Wallner | 1*** | 2/1***/1** | 2/1***/1** |
| Guerois | 0 | 2** | 2** |
| Huang | 0 | 2** | 2** |
| Scorer | Top-1 | Top-5 | Top-10 |
| Karaca | 1** | 2/1***/1** | 2*** |
| HDOCK | 0 | 2/1***/1** | 3/1***/1** |
| Kihara | 0 | 1*** | 2/1***/1** |
Target T312
Only one predictor and scorer group (the same one) managed to submit acceptable models for this target: Kmiecik. A few groups did submit acceptable models in their set of 100 models, but they were not ranked high enough, these were Engel (Haddock team), Guerois, Zou and MDOCKPP.
Target T314
This target was generally well predicted, with in their top-5 acceptable models for Gray, Vakser, and PYDOCKWEB, medium-quality for Huang, HDOCK, Guerois and CLUSPRO, and high-quality models for Wallner. Four scorers managed to produce high-quality models (top-10): Olechnovic, Kihara, Karaca and HDOCK.
Target T315
Also T315 was well predicted, with most predictor groups producing medium-quality models in their top-5 set: Wallner, Brysbaert, Zou, Huang, Guerois, Pierce and Gray. In addition, CLUSPRO produces high-quality models. For scorers (top-10) the picture was similar, with acceptable for Xiaotong Xu, medium for Venclovas, LZERD, Kihara, Zou, MDOCKPP and HDOCK, and high quality for Karaca.
Target T316
No predictor group produced a model of acceptable quality or better in their top-10 submission. However, the shuffled set did contain acceptable models, produced by Pierce, Guerois and PYDOCKWEB. Scorers Shen and HDOCK both recognized a different one of these acceptable models.
Rounds 59 and 60
To follow…
CASP16 - CAPRI Rounds 57 and 58
Round 57 corresponds to the classic joint CASP-CAPRI blind prediction experiment, where only sequence and stoichiometry information is provided. The Round contained 34 targets, which led to a total of 47 assessment units (AUs). Round 58 included a subset of the same targets, for which roughly 8000 MassiveFold models were provided, please see their abstract for details. The MassiveFold models were also included in the Scoring of Round 57. Round 58 had no Scoring Round.
The CASP monomer and multimer categories were merged this year. Marc Lensink’s presentation at the CASP meeting can be found HERE. CASP assessors Nick Grishin and Qian Qong focused on monomeric structure quality and monomeric predictions in the context of multimeric structures. Their presentations will be linked to once available.
Bear in mind that data listed here is subject to minor changes.
Round 57
The assessment results of the top-5 submitted models for Predictors, Scorers, and CASP participants for all individual target interfaces are collected in this file.
The CAPRI ranking, shown in the image below, shows incremental improvement with respect to the baselines provided by COLABFOLD and AlphaFold3 (a manual submission made by the Elofsson team). Highlighted names (Kihara, Cheng, Kozakov/Vajda, Huang, HDOCK, Zou) were predictors who achieved acceptable quality or better for 35 or more of the AUs. Servers are shown in boldface all-caps.
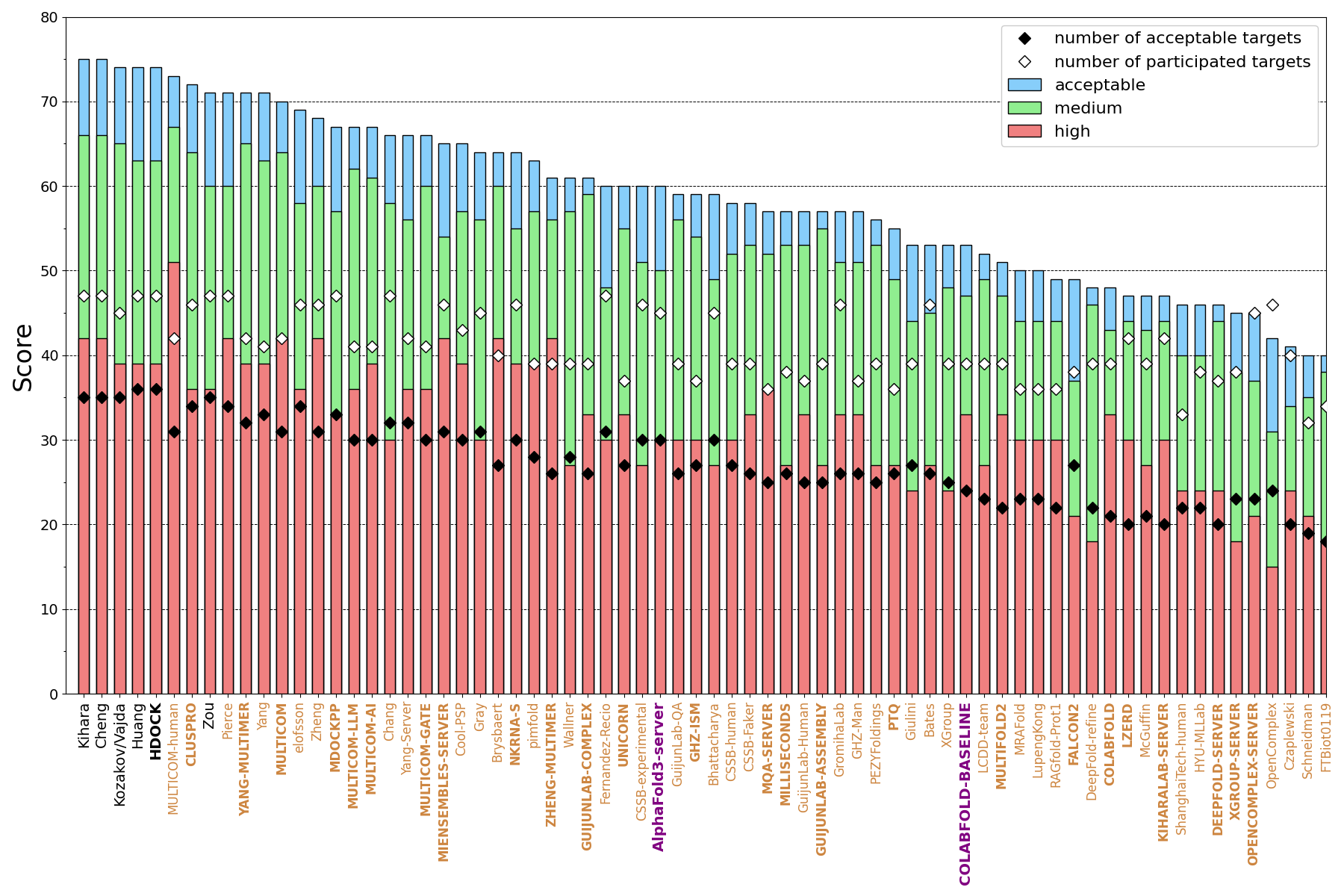
As common for joint CASP-CAPRI Rounds, a ranking based on DockQ Z-scores was also calculated. Z-scores were capped at 2, and negative values were ignored. Such a ranking highlights those participants that performed significantly better for the more difficult targets. The ranking below includes Predictors, Servers, and Scorers. Following top predictor Kozakov/Vajda, you find Yang (and server), scorer Olechnovic, server CLUSPRO, and predictor Kihara.
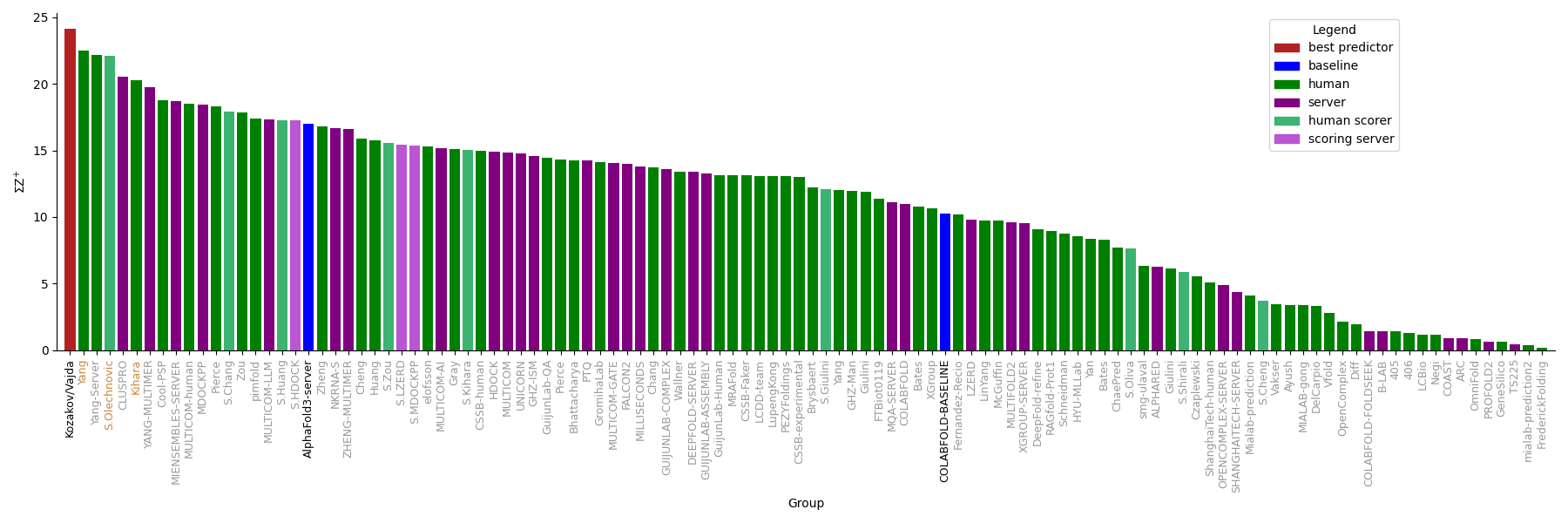
Round 58
Comparing Rounds 57 and 58 on the common set of targets highlights the prediction improvement upon availability of the MassiveFold models. The following graph is ordered on improvement in score, requires a minimum participation of 20 AUs, and a difference in participation for both Rounds of no more than 4 AUs. The assessment results for Round 58 can be found here.
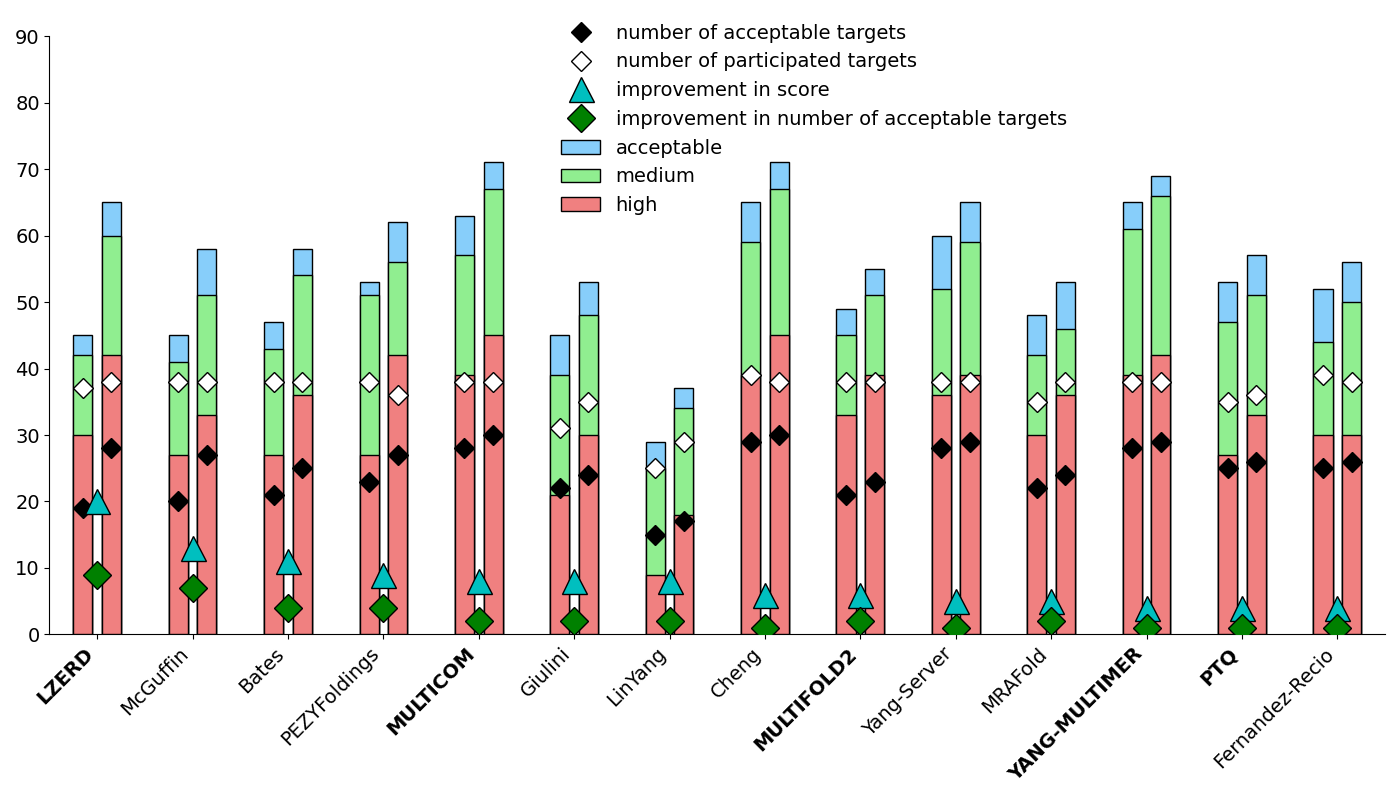
Round 56
Round 56 consisted of one target, an MHC/antibody complex. Four servers (CLUSPRO, HADDOCK, LZERD, MDOCKPP) and another six human predictor groups (Kihara, Zou, Giulini, Brysbaert, Shen) produced medium-quality models in their top-5 submission. Two groups stand out: Kozakov, for producing a high-quality model in top-5, and HADDOCK, for producing a medium-quality model in top-1.
Nine scorer groups including three servers produced medium-quality models in top-5/top-10, of which five in top-1 (Chang, Kihara, Zou, LZERD, MDOCKPP).
The assessment of all models can be found here.
Round 55
CAPRI Round 55 had four targets, three of them with antibodies, including one (T231) binding a peptide. The remaining target (T232) was an EM structure of an existing X-ray structure, however with a different binding mode. For all four targets, the Brysbaert group in collaboration with the assessors had made AlphaFold2 models available, which were also assessed.
The results for all four targets of Round 55 can be found in this comma-separated file.
Target T231
The 25 AlphaFold models featured 22 acceptable quality models, including the first 10 models, 2 medium quality models with models ranked 13 and 14, and one incorrect model. Seven predictor groups (Kozakov/Vajda, Pierce, Karaca, Huang, Venclovas, Zou, Vakser) and 3 servers (CLUSPRO, LZERD, HDOCK) produced medium-quality models in their top-5 submission. An additional 7 predictor groups (Bates, Kihara, S_Chang, Fernandez-Recio, Brysbaert, Giulini, Furman) and one server (PYDOCKWEB) produced acceptable models. Scorer groups Giulini, Huang, Zou and server HDOCK recognized medium quality models, while scorer groups Kihara, Bates, Venclovas, Karaca and server LZERD recognized acceptable models.
Target T232
The 25 AlphaFold models were all of medium quality. More than half of the predictor groups and 80% of the scorer groups also produced 5/5** models. With a few rare exceptions, there were no models of acceptable quality.
Targets T233 and T234
These two targets featured binding of two different Fab’s to a major histocompatibility complex-I. All AlphaFold models were incorrect. Interestingly, a few groups managed medium-quality solutions (and no acceptable!) for T233. These were S_Chang, Kihara, Zou, Venclovas, Huang and server HDOCK, which were recognized by scorer groups Venclovas and Zou. For T234, no acceptable models were submitted by predictors, but a few acceptable and one medium quality model were available in the shuffled set. Only the Giulini scorer group recognized an acceptable quality model, and submitted it as their scorer model 10.
Round 54 CASP15-CAPRI
CAPRI Round 54 comprised 37 targets, divided into 38 Assessment Units (AU). The assessment results of the top-5 submitted models for Predictors, Scorers, and CASP Assembly Prediction participants for all individual target interfaces are collected in this comma-separated file. For explanation of the invididual columns we refer to the CAPRI Scoreset v2022 help page.
Most AUs are single-target single-interface, with the exception of the following:
| Target | AU 1 | AU 2 |
|---|---|---|
| T203 | average of interfaces 1-3 | interface 4 |
| T204 | best of interfaces 1-2 | average of interfaces 3-8 |
| T219/T220/T221 | best of interfaces 1-3 | best of interfaces 4 |
You can find the slides of Marc Lensink’s CAPRI presentation in the CASP assembly session HERE.
The image below shows the CAPRI ranking. “AF2-MULTIMER” is the off-the-bench AlphaFold-Multimer modeling as submitted by the Elofsson group.
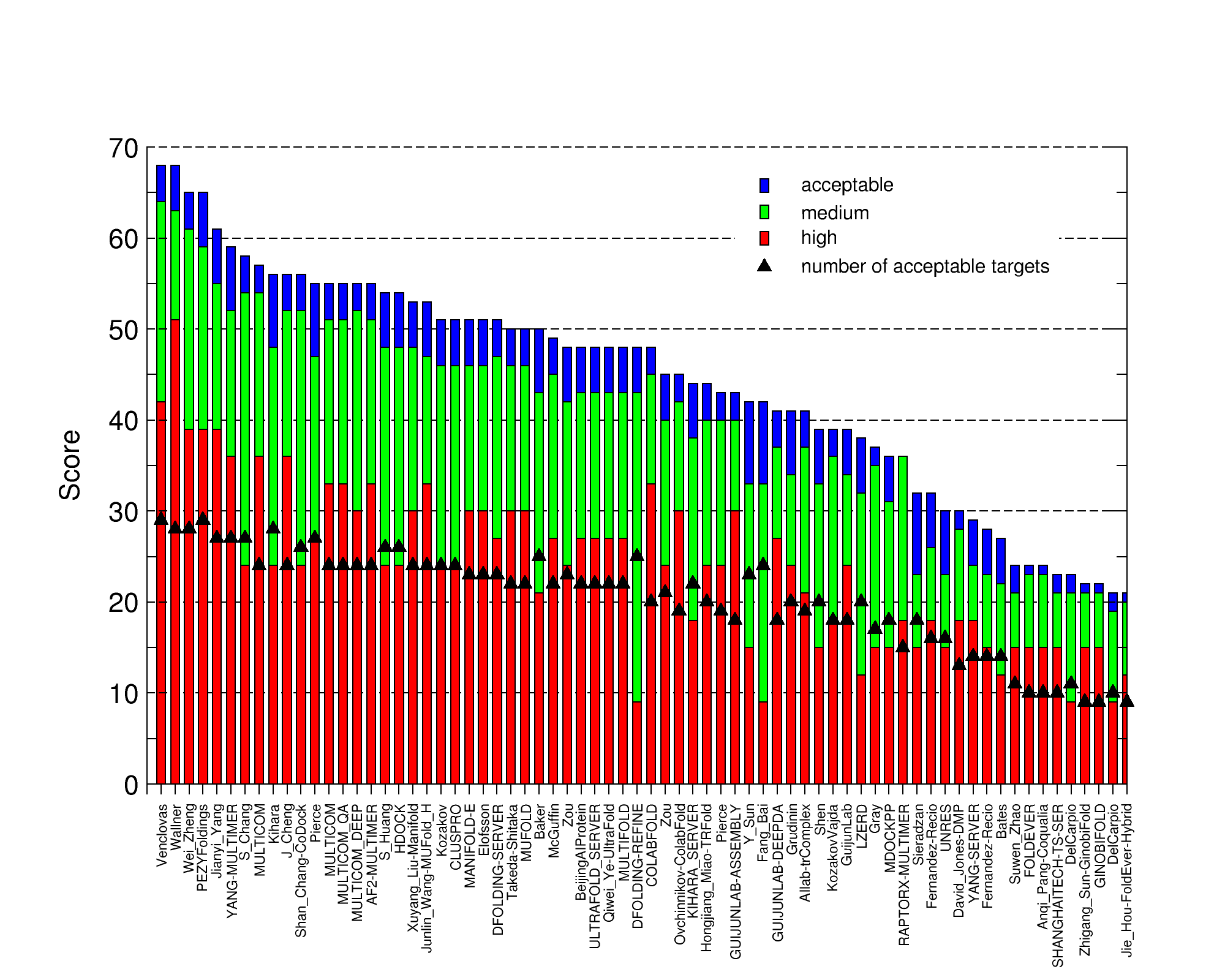
Round 53 Target T187 and T188
Targets T187 and T188 featured a large conformational change upon binding to dsDNA. However, the AlphaFold monomer structure captured this conformational change perfectly. Unfortunately, no predictors or scorers were able to produce acceptable models for T187. The assessment results for the top-5 submitted models are collected in this comma-separated file. In this file, interface 1 corresponds to the whole target; interface 2 is the protein dimer alone.
For T188 the bound dsDNA structure was given. This resulted in acceptable models for 2 predictor groups: Fernandez-Recio (top-1) and Pierce (top-5); and 5 scorer groups: Fernandez-Recio, Zou and server PYDOCKDNAWEB (top-1), and Kihara and server MDOCKPP (top-5).
Medium-quality models for the protein dimer were produced (in top-5 submission) by Venclovas, Zacharias, Huang, Chang, Kihara, and servers LZERD and HDOCK. Scorers Kihara, Bates, Huang, Chang, Zou, Shen, and servers HDOCK, LZERD and PYDOCKDNAWEB recognized these (again, top-5).
The quality of the protein-dnDNA models strongly depended on the protein dimer modelling quality, as can be seen in the image below.
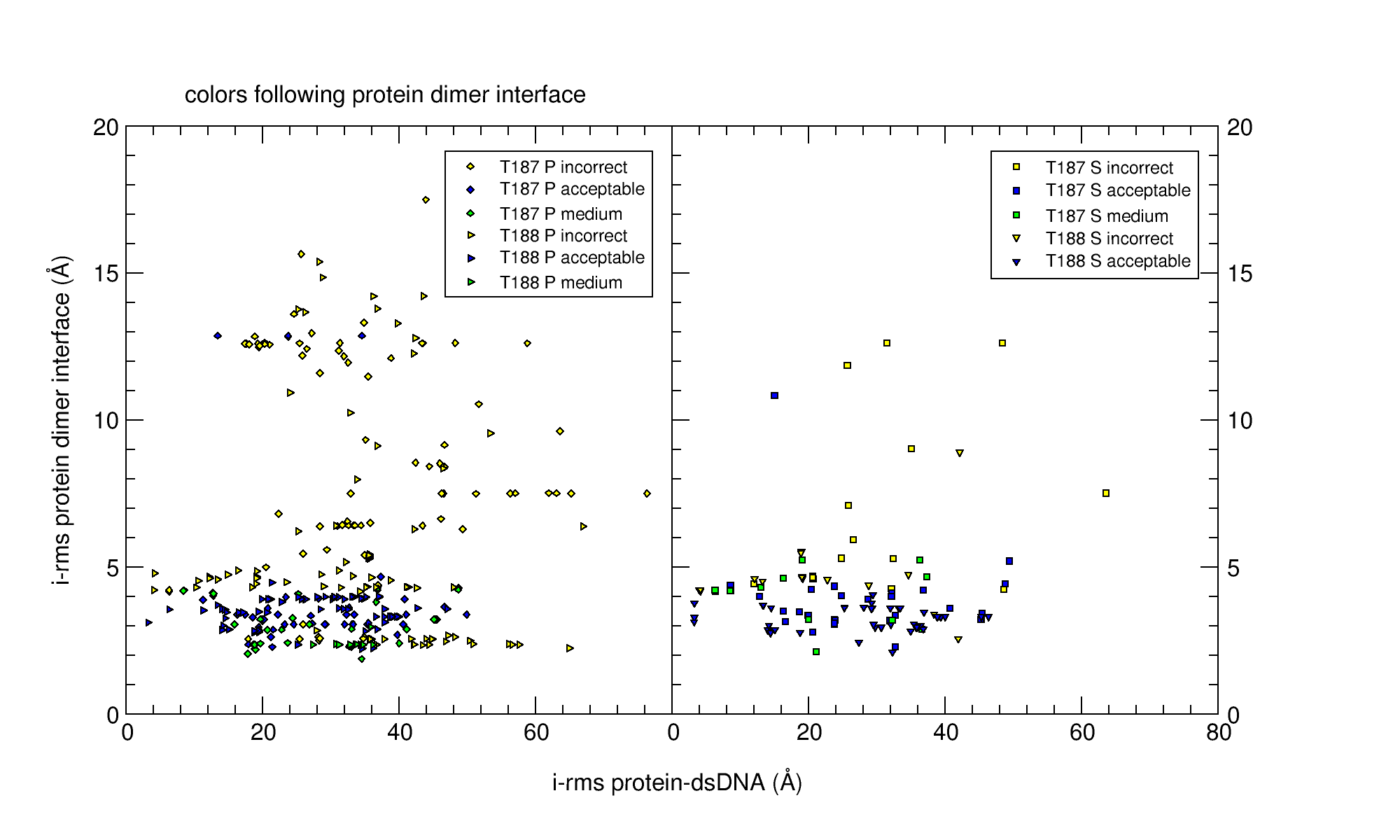
Round 52 Target 186
The challenge of T186 was to predict the conformation of a ~100 residue loop lying on the surface of a large pocket. The loop was well structured but without any secondary structure elements.
Close to 25% of the submitted predictions were missing any coordinates for the loop residues and were therefore disqualified. Close to half of these were of medium quality (for the structure without the loop), as a high quality template was available for the rest of the protein assembly.
The loop was part of chain G and in contact with the rest of the chain as well as chain D. Three assessment were performed: (1) between chains G and D, (2) between the loop and the rest of chain G, and (3) between the loop and chain D. Only acceptable quality models were obtained.
Predictor groups Kihara and Scorer groups Kihara and Venclovas obtained acceptable models for both T186.1 as well as T186.2. Predictor groups Chang, Furman, Fernandez-Recio, Huang and Scorer groups Oliva, Chang, LZERD, HDOCK, Huang, Zou, MDOCKPP, Fernandez-Recio, Shen, J_Huang only for T186.1. No Predictor nor Scorer groups produced acceptable models for T186.3.
The full list of assessment is here, do note that this list includes the disqualified models (look for the “clashes” or “low_id” flag in columns 36 and 37, resp.
Round 49 Target T163
Target T163 features a 2:2 hetero-tetramer, with 2 copies of SYCE2 and 2 copies of TEX12. It has now been published as PDB 6R17. T163 contains 2 homo-dimeric interfaces: T163.1 (SYCE2) and T163.5 (TEX12); and 3 hetero-dimeric interfaces: T163.2 (SYCE2.Nt/TEX12), T163.3 (SYCE2’.Ct/TEX12) and T163.4 (SYCE2.Ct/TEX12’).
A homo-dimeric structure of TEX12 existed, but it did not correspond to the binding mode in T163. This might have complicated the prediction of this target. The assessment results for the 5 interfaces can be found in this comma-separated file.
Predictors and Scorers with models of acceptable quality or better in their top-5 submitted models are listed in the following table:
| Interface | Predictors | Scorers |
|---|---|---|
| T163.1 | - | Fernandez-Recio |
| T163.5 | Kihara (medium) | Kihara (medium) |
| T163.2 | Venclovas, Kihara, Gray | Seok (medium), Oliva, Perthold |
| T163.3 | - | - |
| T163.4 | LZERD | - |
Round 48 Targets T161 and T162
Both targets feature the same toxin-antitoxin complex, a hetero-complex consisting of a toxin homo-dimer bound to two antitoxin molecules. The challenge of T161 was to predict the bound form of the toxin homodimer, which forms a completely different structure than in its unbound form. The challenge for T162 was to predict the structure of the full toxin-antitoxin complex. For T162, the bound structure of the antitoxin was supplied, with side-chains removed, as well as all submitted models (predictors and scorers) of the toxin homo-dimer (T162). The full assessment is here.
Only two groups managed to produce good models for these targets:
| Target | Group | Performance |
|---|---|---|
| T161 | Bates | 1 |
| T162 | Andreani/Guerois | 2/1** |
Round 47 Target T160
Target T160 features the structure of the Bacillus anthracis Sap S-layer assembly domain, now published as PDB 6HHU. It consists of 6 individual domains that are organized in two dimensions. To it are bound two nanobodies. The individual interfaces bury between 150 and 610 Å2. The interfaces are indicated in the following picture of the target.
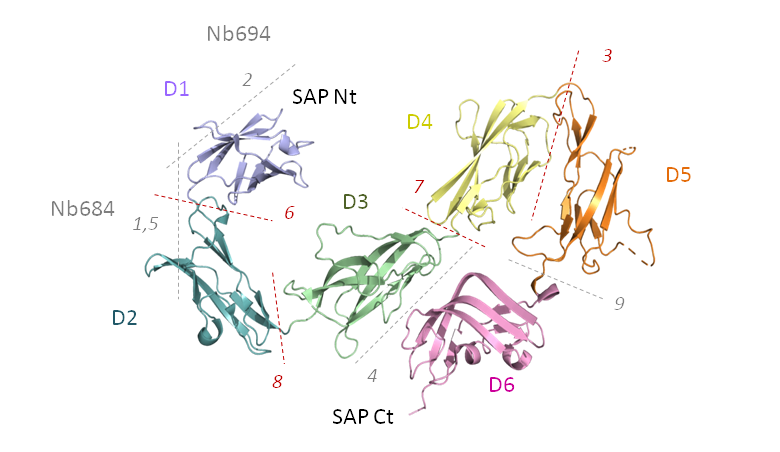
Nanobody Nb694 is bound to domain D1 through interface T160.2, while Nb684 is bound to domains D1 and D2 through interfaces T160.1 and T160.5. Whereas the interface between domains 1 and 2 was best predicted, nobody managed to identify the nanobody binding modes. The results for all interfaces can be found in this comma-separated file. Predictors and Scorers with models of acceptable quality or better in their top-5 submitted models are listed in the following table:
| Interface | Predictors | Scorers |
|---|---|---|
| T160.2 | - | - |
| T160.1 | - | - |
| T160.5 | - | - |
| T160.6 | GALAXYPPDOCK, Fernandez-Recio, | Fernandez-Recio, Kihara, |
| Andreani/Guerois | LZERD, Venclovas, Bates | |
| T160.8 | Andreani/Guerois (medium), | Venclovas (medium) |
| Kozakov/Vajda (medium), Venclovas (medium) | ||
| T160.7 | Gray | - |
| T160.3 | CLUSPRO (medium), Kozakov/Vajda (medium), | Venclovas (medium) |
| Andreani/Guerois (medium), Venclovas (medium) | ||
| T160.9 | - | HDOCK |
| T160.4 | - | - |
For questions or comments please contact Marc Lensink

